OpenEye Cup 2025

Talk Notes#
DELs#
-
Tango:
- crisp cas for synthetic lethality
- immune evasion
- “cold vs hot”
- you need ligands for your targets.
- DEL for Hit Generation
- Idea: is there a good Rust-based DEL debarcoding library?
- 50 libraries 3B compounds.
- Binding Enrichment
- how do they compare against docking.
- how many mols do you need?
- enzymatic activity?
- Binding?
- QPCR between steps.
-
Michael Hack from J&J
- comm chem
- pool-split-pool
- “DEL Cube” - looking for activity on “lines”, “planes”, and “spots”
- where planes represent an active pharmacophore with potential SAR activity already built in.
- DEL targets are starting points.
- typically verified “off DNA”
- DB size if ~16TB
- Cephalogix
- Triage via Gini Coefficient.
-
James Wellnitz:
- structure genomics consortium
- DEL affinity data
- aim to have specific probes for each protein.
- Major shade on the capability of DEL data to extend beyond the tested interaction.
- Very low predictability of 1 DEL dataset models compared to others.
-
Honyuan Zhu:
- Design and analysis of DEL experiments
- Pfizer ML & CS
- 1000x1000x100 compounds == 1B
- created dataset so 3x1000x100 = 3M which is more manageable
Ivet Behar#
- elastic network models
- Analytical not simulation
- anisotropic network modeling
- “Hessian”
- Target Hinge sites within proteins.
- Weighted Ensembled for fast MD approximation
- Rhapsody for mutational effects
- Elastic Networks to sample conformational space for use with WE
David LeBard#
- Use of normal mode analysis
- use Xenon as a probe
- identify and score pockets for their ability to generate hydrophobic pockets.
Ken Dill#
- opening joke: scientists DNA, PRoteins, RNA scientist “get out my chair”
- Multiple Potential origin of life stories:
- RNA world; lipid world, metabolism first, other
- Focus not on the immediate origin but how do you “escape entropy” - e. g how do transition from a deterministic entropy-driven system to one that can make more complex components against entropy:
- “to get from chemistry to biology you need you need cooperativity”
- presented an analogy of Origin of life to protein folding problem:
- landscape seems improbably big but it is an energetic funnel.
- what would be the effect of funnels on origins?
- laid out a simple model of hydrophic vs hydrophilic polymers self organizing
- cna these folded bits organize and enhance generation of more of themseleves?
- e.g. can they be autocatalytic? can they cooperatively assist?
- Ken shows highlights of decades work on peptoids showing:
- random mixes of hydrophic/polar peptoid chains will fold
- folded bits lead to longer folded bits providing evidence for auto catalysis
- He concludes with the close that:
- none of the other competing theories for life can explain the transition from “soup of chemicals” to “cooperative autocatalyic mechanism”
- his mechanism of simple physical forces acting on diverse polymers can provide such a mechanism.
- he belives his peptoid work shows that “foldcats” do actually act as incredibly basic but working catalytic mechansims.
- that early transition to biological systems form chemistry must have been mediated by proteins.
Michael Levine#
- AI for physics in science
- Head of Comp Chem at Genesis
- “anyone who tells you they have an end-to-end AI discovery engine is …..”
- AI tool sup and down the stack but with strong human intervention and curation at all stes.
- Hybrid Systems work well.
- AI systems: Interpretion vs Extrapolation
- Discussion of limitations of pure co-folding models (brought home later by P. Walters)
- “we are deciduos”
- In regards to if a generative model is any good I run a process called “Med Chem Tinder”
- Structures can suck - many example sof misplaced Densities etc.
- “can we retain models on raw density….” Someone… can
Pat Walters#
- Opening slide: “Warning: Rant Ahead”
- Begins with caveat that he is ALL ABOUT using computational tools including AI for Drug Discovert. Her even got his PhD in expert systems. However…..
- A major launch against the AI hype cycle in Drug Discovery and where the various modeling tools fall apart.
- He enumerates a number of papers, blog posts, and press releases and then systematically exposes the fundamental weaknesses of the methods and how, once the weakness is exposed, the claims basically fall apart.
- Point 1: Many “AI designed” molecules are trivial modifications to existing compounds working on existing targets - in other words there is basically no value added as the molecule is essentially the same.
- Point 2: Publications of Untested Generated Compounds Produce Really Bad Molecules. He chose a specific example and showed that of the 1000’s of compounds published in one paper, the majority (>90%) contained ring systems that have never been characterized and are fundamentally chemically unstable.
- Point 3: LLMs are great at syntax but NOT Chemistry. He showed a series of examples using LLMs to create modifications of a starting SMILES string or IUPAC identifier and analyzed the outputs. There are many valid SMILES strings/ IUPAC strings that are spit out. However, he shows how many of these are either trivial, non-syntheziable, or completely chemically wrong.
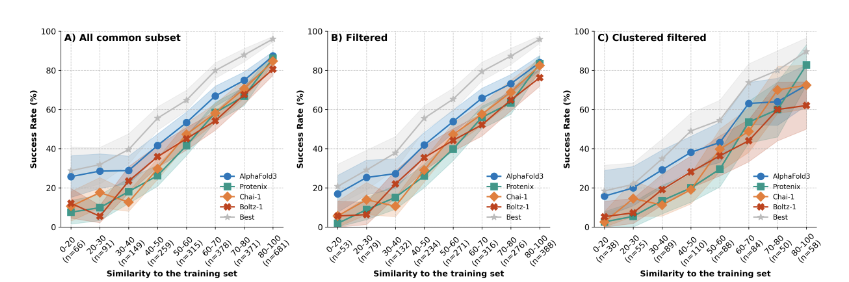
- Point 4: Reports of Model Efficacy are largely wrong because they don’t control for memorization in the dataset. New “Must Read” paper: Have Protein-Ligand Co-Folding Methods Moved Beyond Memorization?
- Summary: Everyone is using LLMs and AI. Its a beautiful wonderful time to be in science and have these tools BUT these are fundamentally tools and have been wildly oversold as a replacement for a long difficult process.
- Very enjoyable.
Other Notes#
-
Claudio Catalano:
- we throw away 90% of Cryo EM data
- 10 mg/ml in Cyo EM is a lot.
- a single 3D classification can take 4-5 days.
- navigating paths between stable conformers is of interest
-
Michael Wall
- use more of the X-ray diffraction data ro understand motion in X-tals.
- mentioned X-ray lasers as a new an upcoming tool
-
Bowen:
- attempts to use subsampled MSAs with ALphaFold to generate new protein conformations.
- use the DB of lots of proteins snap,es at 300ns of MD
- Atlas Database
- “root mean squared wasserstein distance” - generalized RMSD
-
Handley:
- jokes on being “out of band
- overview of genomic language models and transformer architecture.
- Shows EVO2 // GeneFormer
-
Jenna Fromer:
- Opens with a hat tip to Anthony Nicholls: “why should a human do the choosing?”
- Introduces Sparrow a constraint solver for automatically choosing molecules that takes into consideration cost, synthetic route, batch size an reusability.
- Got a number of questions from the audience about how that would work on their particular campaigns and Jenna notes that there are tweaks under way to address these cases.
CUP Schedule#
CUP is OpenEye’s annual scientific meeting held at La Fonda on the Plaza in Santa Fe where we bring together top Scientists, Customers, Users, and Programmers in the industry to discuss the challenges in drug discovery. The event features reports from OpenEye scientists, two keynote speakers and four half-day sessions.
Keynote Speakers#
- Frank K. Brown Industry Perspective: Jeff Blaney - Sr. Director, Discovery Chemistry, Genentech
- The Levinthal Lecture: Ken Dill - Laufer Family Endowed Professor, Stony Brook University
Sessions#
Monday PM: Santa Fe Room
- 7:00 - Welcome Reception
Tuesday AM: Lumpkins Ballroom - OpenEye: Scientific and Platform Advances
- 8:00 - Registration Opens - Breakfast buffet on the Mezzanine
- 9:00 - What’s Going on Around Here? - Geoff Skillman, Vice President of R&D, BU Head
- 9:30 - Just When You Thought We had Reached the Search Limit - Matt Geballe, Group Director, Scientific R&D
- 10:00 - OMEGA is the Alpha & Bayes - Riddhish Pandharkar, Scientific Developer
- 10:30 - Break
- 11:00 - Dude, Can I Bind Here? - Neha Vithani, Scientific Developer
- 11:30 - There Were Always Multiple Conformations! - David Wych, Scientific Developer
- 12:00 - Lunch Break (on own)
Tuesday PM: Lumpkins Ballroom - OpenEye: Scientific and Platform Advances
- 2:00 - AI AI Everywhere, AI AI In My… - Caitlin Bannan, Manager, Scientific R&D and Jesper Sørensen, Head of Scientific Development
- 2:20 - Orion is Not One-Size Fits All - Jharrod LaFon, Vice President of R&D
- 2:40 - The Haystack is Massive - but We Find the Needles Fast - Kalistyn Burley, Application Scientist
- 3:00 - Break
- 3:30 - AI Looks Better With 3D Colored Glasses – Jingyi Chen, Scientific Developer
- 4:00 - A Map Guiding You to the Right Trees in the Forest - Chris Neale, Senior Manager, Scientific R&D
- 4:30 - Minimizing Your Losses…Lessons from Rhode Island Hold-Em - Greg Bakken, Group Director, Scientific R&D
- 5:00 - Break
- 5:15 - Frank K. Brown Industry Perspective Keynote: “What I’ve Learned from 42 Years of Drug Discovery: Douglas Adams was Right - Jeff Blaney - Sr. Director, Discovery Chemistry, Genentech
- 6:30 - Reception & Dinner Buffet - La Terraza Banquet Room
Wednesday AM: Lumpkins Ballroom - Computational DEL Analysis
- 8:00 - Registration Opens - Breakfast buffet on the Mezzanine
- 9:00 - Ligand Discovery Using DEL Screening - William Mallender - Vice President of Biochemistry, Tango Therapeutics
- 9:30 - Using computation to design and explore compound libraries, with an emphasis on DELs - David Mobley - Professor, UCI Department of Chemistry
- 10:00 - Break
- 10:30 - J&J’s DEL Informatics Platform: Perspectives from a (Relative) Newcomer – Michael Hack - Senior Principal Scientist, Janssen Pharmaceuticals
- 11:00 - The potential, and pitfalls, of DEL for big data generation - James Wellnitz - Tropsha and Popov Groups, Dept. of Chemical Biology and Med. Chem., UNC
- 11:30 - Computational approaches enabling PF-DEL platform: from library design to target screening - Hongyao Zhu - Senior Principal Scientist, Pfizer Inc.
- 12:00 - Lunch Break (on own)
Wednesday PM: Lumpkins Ballroom - Cryptic Pockets and Allostery in Drug Discovery
- 2:00 - Computational Assessment of Positive & Negative Allosteric Modulators of GPCRs - Ivet Bahar - Louis and Beatrice Laufer Endowed Chair, Professor, Department of Biochemistry and Cell Biology, Stony Brook University
- 2:30 - Rapid Molecular Modeling for Focused Cryptic Pocket Identification - Yunhui Ge - Scientist, Alkermes
- 3:00 - Illuminating an invisible HIV-1 capsid protein state via 19F NMR and weighted ensemble simulations - Lillian Chong - Professor, Department of Chemistry, University of Pittsburgh
- 3:30 - Break
- 4:00 - There’s Plenty of Room in Your Undruggable Protein - David LeBard - Head of Science, OpenEye, Cadence Molecular Sciences
- 4:45 - Break
- 5:00 - The Levinthal Lecture: Ken Dill - Laufer Family Endowed Professor, Stony Brook University
- 6:00 - Poster Session with Dinner Buffet - New Mexico and Santa Fe Ballrooms
Thursday AM: Lumpkins Ballroom - Modern Techniques in Structural Biology
- 8:00 - Registration Opens - Breakfast buffet on the Mezzanine
- 9:00 - Reconstructing Conformational States & Densities in CryoEM with RECOVAR – Marc Aurèle T. Gilles - Assistant Professor, Department of Mathematics, Princeton University
- 9:30 - Bridging the Gap: Unlocking the Potential of CryoEM for Dynamic Ensembles – Claudio Catalano, Nanoimaging Services
- 10:00 - Break
- 10:30 - CryoDRGN: Advances in Deep Learning for Reconstructing Protein Structure & Dynamics– Rishwanth Raghu - PhD Candidate, Zhong Lab, Princeton University
- 11:15 - Molecular-Dynamics Simulations for Protein Crystallography – Michael Wall - Senior Scientist, Los Alamos National Laboratory
- 12:00 - Lunch Break (on own)
Thursday PM: Lumpkins Ballroom - Separating the Wheat from the Chaff: Where and When AI Works in Drug Discovery
- 2:00 - GEMS: Combing AI and Physics To Advance Small Molecule Drug Discovery – Michael LeVine - Head of Computational Chemistry, Director of Computational Biophysics, Genesis Therapeutics
- 2:30 - Towards Emulation of Molecular Dynamics with Generative Deep Learning – Bowen Jing - PhD Candidate, Berger Group, CSAIL, MIT
- 3:00 - Break
- 3:30 - An Introduction to Modern Genomic Foundation Models - Simon Handley - Sr. AI/ML Solutions Architect, AWS Healthcare and Life Sciences
- 4:00 - Balancing Utility & Synthesizability for Actionable AI-driven Molecular Design – Jenna Fromer - PhD Candidate, Coley Group, MIT
- 4:30 - Artificial Intelligence in Drug Discovery – Revolution, Evolution, or Complete Nonsense – Pat Walters - Chief Data Officer, Relay Therapeutics
- 5:00 - Closing Remarks - Geoff Skillman, Vice President of R&D, BU Head
- 6:00 - Conference Dinner at Casa España, 321 W San Francisco St. Santa Fe, NM
Resource Links#
- Cephalogix
- Structural Genomics Consortium
- Leash Bio
- HitGen
- OpenEye
- Orion Platform
- ROCS
- Schrödinger
- University of Pittsburgh Chemistry Department
- WESTPA Overview
- WESTPA GitHub Repository
- Presentation Slides
- ProDy GitHub Repository
- Rhapsody
- Peptoid - Wikipedia
- Royal Society Publishing Article
- NanoImaging Services
- LANL GitHub
- ATLAS Database
- ChEMBL Database
- Sparrow GitHub Repository
- bioRxiv Preprint
Some Santa Fe Pics#